An optokinetic tool for in vivo protein expression control is used for diabetes therapy

Kristine Lyxa
Research sceintist @ librrar
Here to bring you recent news on our vivo protein exposure
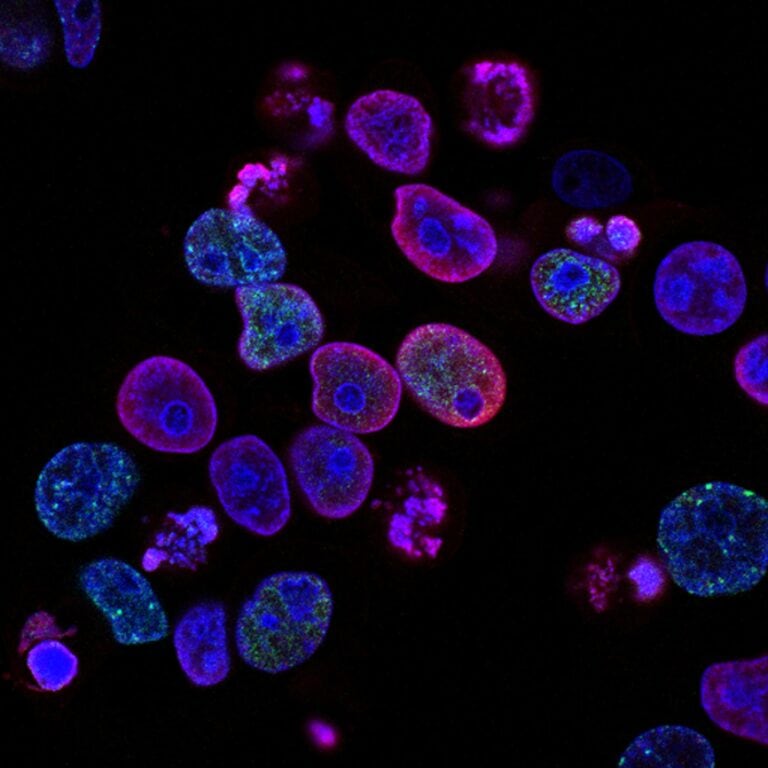
Chinese scientists have created an optogenetic tool that can accurately regulate gene expression by mimicking normal expression in living systems. The scheme presented by them will help to study in depth the mechanisms of gene expression in animal cells. In a paper published in Nature Communications, the researchers also demonstrated the potential use of the molecular tool as a therapy for type 1 diabetes.
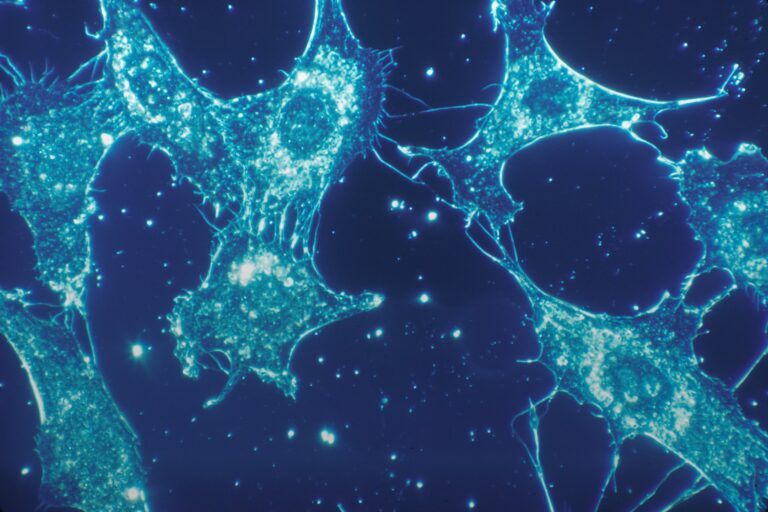
To understand how a particular protein functions, how a cell responds to external and internal signals, it is necessary to be able to analyze how genes are expressed, and, accordingly, proteins are produced. It is generally accepted that under constant external conditions, the concentration and functions of key regulatory molecules in the cell do not change much or randomly fluctuate around a fixed value.
However, many regulatory proteins, including transcription factors, function in a “pulsating” manner and, accordingly, differently affect the cell fate, the cell’s response to stress conditions and differentiation. Scientists have managed to explain the mechanisms of only some of these pulsating mechanisms. The creation of controlled pulsating systems can be useful for further study of their mechanism and the biological functions behind them.
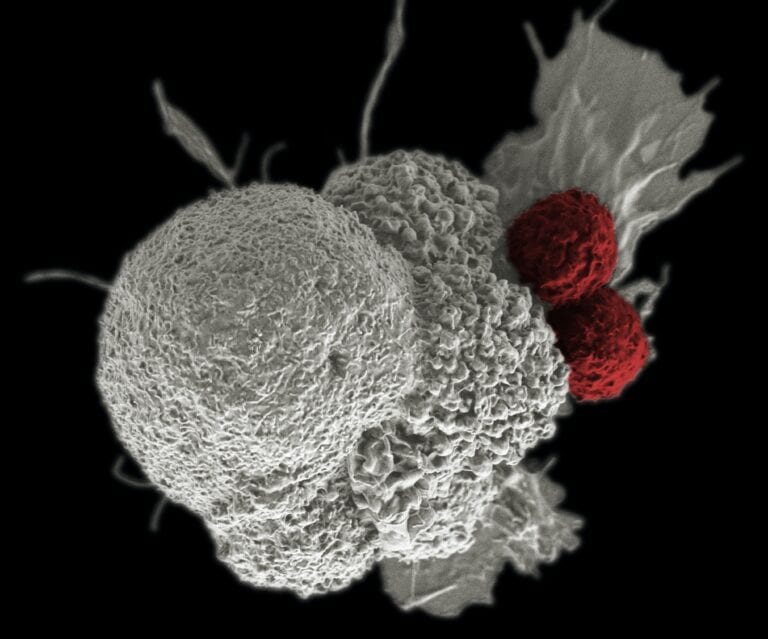
The release of insulin by cells refers to this pulsating pattern. Insulin is a protein hormone secreted by the beta cells of the pancreas. Insulin blocks the release of glucose from the liver and encourages cells to use glucose from food in the blood.
In research work, chemicals that influence gene expression are widely used to control gene expression. However, such chemical inducers do not disintegrate for a long time and therefore affect expression for too long. And if in an experiment on cell cultures it is possible to physically remove the acting substance (change the environment), then in animal experiments there is no such possibility (it is impossible to “take back” a substance from animal cells), and it is practically impossible to dynamically regulate expression by simply adding or removing a chemical … There are alternative systems in which gene expression depends on light, but the use of such systems is very limited: it is difficult to “deliver” the required amount of light deep into the tissue, because the tissue itself absorbs light quite strongly.
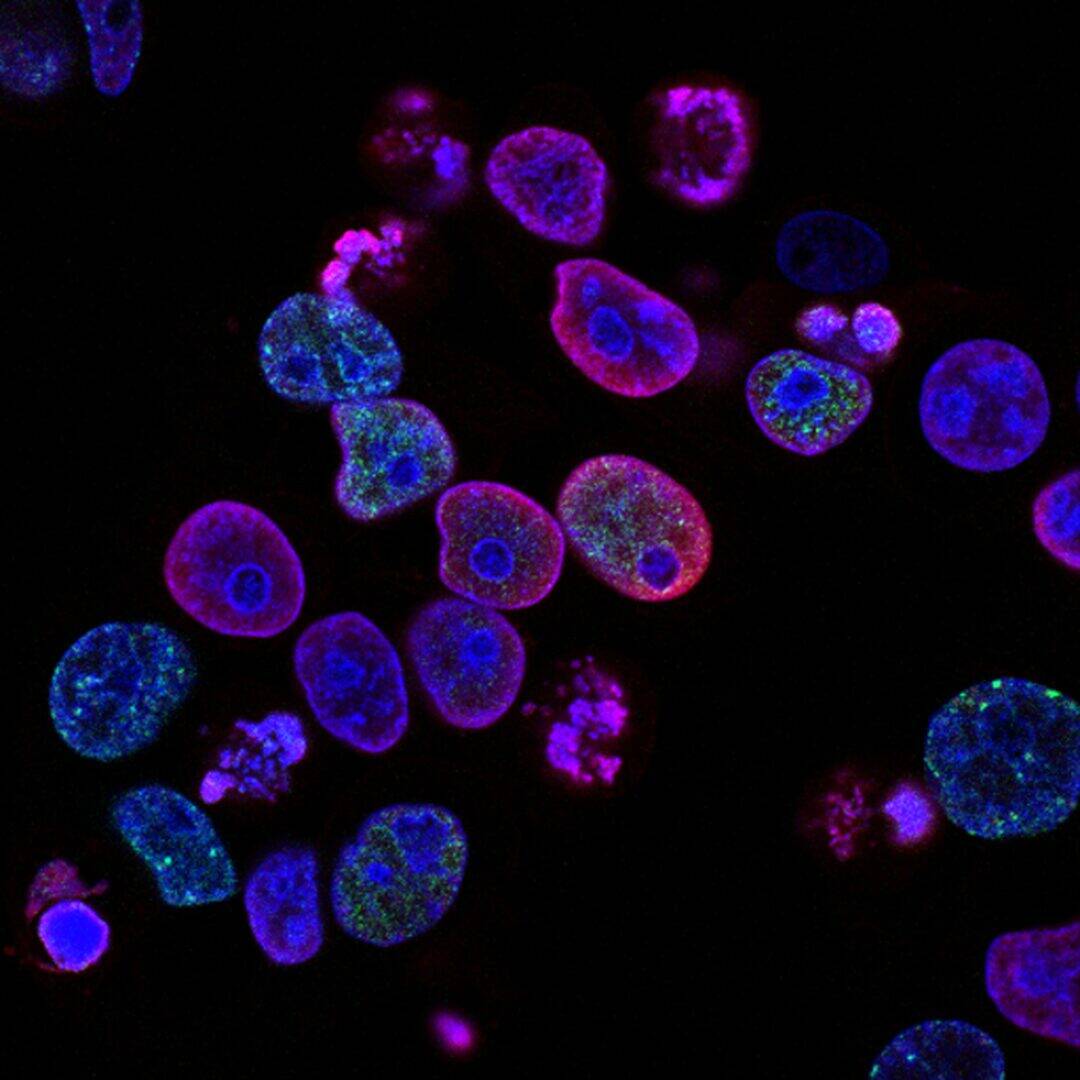
The problem of protein management is of interest not only to those involved in gene expression. Previously, neuroscientists had developed chimeric proteins that simultaneously possess both luminescence (the ability to glow) and light sensitivity.
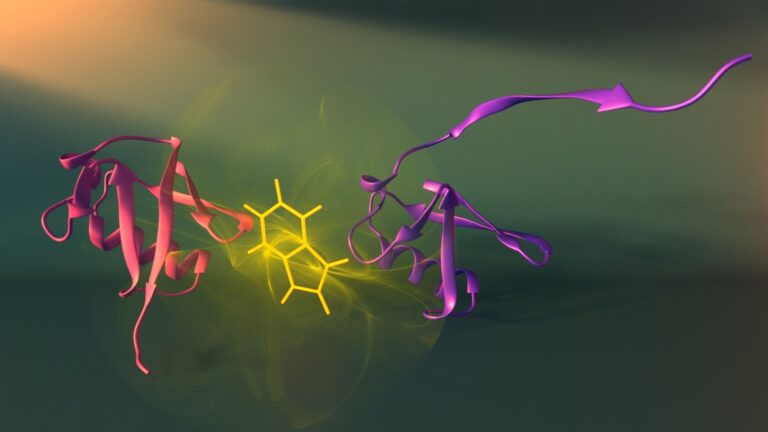
A group of scientists from the East China University of Science and Technology, led by Yi Yang, proposed using a similar system, which combines chemical and light regulation, to control gene expression in cells. The scientists decided to create a chimeric protein consisting of a light-activated transcription factor and luciferase. To do this, they combined a modified deep-sea shrimp luciferase NanoLuc and the LOV domain (light-oxygen-voltage), a light-sensitive protein of plants and fungi, into one molecule. In this case, the LOV domain absorbs blue light well (wavelength λ = 440–480 nanometers), and the NLuc luciferase just produces blue light. This system can be influenced in two ways: directly by light or by providing luciferase with its substrate, furimazine. Experiments have shown that the system actually functions as a logical “OR” element. The expression of the protein controlled by the chimeric transcription factor increased 116-fold with the addition of furimazine at a concentration of 2.5 micromolar, and decreased with a further increase in the substrate concentration.